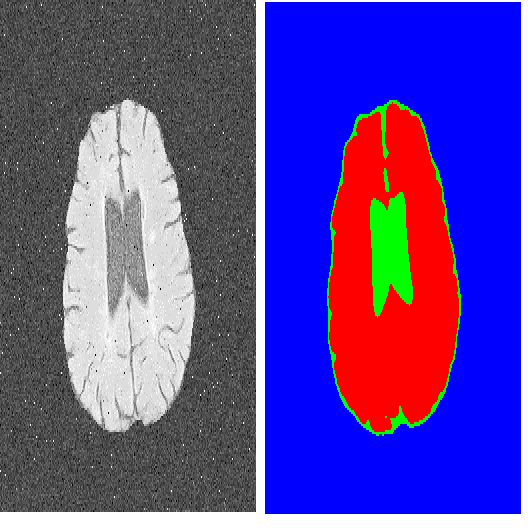
Deep learning-based semi-supervised learning (SSL) algorithms have led to promising results in medical images segmentation and can alleviate doctors' expensive annotations by leveraging unlabeled data. However, most of the existing SSL algorithms in literature tend to regularize the model training by perturbing networks and/or data. Observing that multi/dual-task learning attends to various levels of information which have inherent prediction perturbation, we ask the question in this work: can we explicitly build task-level regularization rather than implicitly constructing networks- and/or data-level perturbation-and-transformation for SSL? To answer this question, we propose a novel dual-task-consistency semi-supervised framework for the first time. Concretely, we use a dual-task deep network that jointly predicts a pixel-wise segmentation map and a geometry-aware level set representation of the target. The level set representation is converted to an approximated segmentation map through a differentiable task transform layer. Simultaneously, we introduce a dual-task consistency regularization between the level set-derived segmentation maps and directly predicted segmentation maps for both labeled and unlabeled data. Extensive experiments on two public datasets show that our method can largely improve the performance by incorporating the unlabeled data. Meanwhile, our framework outperforms the state-of-the-art semi-supervised medical image segmentation methods. Code is available at: //github.com/Luoxd1996/DTC
相關內容
A key requirement for the success of supervised deep learning is a large labeled dataset - a condition that is difficult to meet in medical image analysis. Self-supervised learning (SSL) can help in this regard by providing a strategy to pre-train a neural network with unlabeled data, followed by fine-tuning for a downstream task with limited annotations. Contrastive learning, a particular variant of SSL, is a powerful technique for learning image-level representations. In this work, we propose strategies for extending the contrastive learning framework for segmentation of volumetric medical images in the semi-supervised setting with limited annotations, by leveraging domain-specific and problem-specific cues. Specifically, we propose (1) novel contrasting strategies that leverage structural similarity across volumetric medical images (domain-specific cue) and (2) a local version of the contrastive loss to learn distinctive representations of local regions that are useful for per-pixel segmentation (problem-specific cue). We carry out an extensive evaluation on three Magnetic Resonance Imaging (MRI) datasets. In the limited annotation setting, the proposed method yields substantial improvements compared to other self-supervision and semi-supervised learning techniques. When combined with a simple data augmentation technique, the proposed method reaches within 8% of benchmark performance using only two labeled MRI volumes for training, corresponding to only 4% (for ACDC) of the training data used to train the benchmark.
Modern neural network training relies heavily on data augmentation for improved generalization. After the initial success of label-preserving augmentations, there has been a recent surge of interest in label-perturbing approaches, which combine features and labels across training samples to smooth the learned decision surface. In this paper, we propose a new augmentation method that leverages the first and second moments extracted and re-injected by feature normalization. We replace the moments of the learned features of one training image by those of another, and also interpolate the target labels. As our approach is fast, operates entirely in feature space, and mixes different signals than prior methods, one can effectively combine it with existing augmentation methods. We demonstrate its efficacy across benchmark data sets in computer vision, speech, and natural language processing, where it consistently improves the generalization performance of highly competitive baseline networks.
In this paper, we aim to improve the performance of semantic image segmentation in a semi-supervised setting in which training is effectuated with a reduced set of annotated images and additional non-annotated images. We present a method based on an ensemble of deep segmentation models. Each model is trained on a subset of the annotated data, and uses the non-annotated images to exchange information with the other models, similar to co-training. Even if each model learns on the same non-annotated images, diversity is preserved with the use of adversarial samples. Our results show that this ability to simultaneously train models, which exchange knowledge while preserving diversity, leads to state-of-the-art results on two challenging medical image datasets.
In this work, we study the problem of training deep networks for semantic image segmentation using only a fraction of annotated images, which may significantly reduce human annotation efforts. Particularly, we propose a strategy that exploits the unpaired image style transfer capabilities of CycleGAN in semi-supervised segmentation. Unlike recent works using adversarial learning for semi-supervised segmentation, we enforce cycle consistency to learn a bidirectional mapping between unpaired images and segmentation masks. This adds an unsupervised regularization effect that boosts the segmentation performance when annotated data is limited. Experiments on three different public segmentation benchmarks (PASCAL VOC 2012, Cityscapes and ACDC) demonstrate the effectiveness of the proposed method. The proposed model achieves 2-4% of improvement with respect to the baseline and outperforms recent approaches for this task, particularly in low labeled data regime.
Despite much success, deep learning generally does not perform well with small labeled training sets. In these scenarios, data augmentation has shown much promise in alleviating the need for more labeled data, but it so far has mostly been applied in supervised settings and achieved limited gains. In this work, we propose to apply data augmentation to unlabeled data in a semi-supervised learning setting. Our method, named Unsupervised Data Augmentation or UDA, encourages the model predictions to be consistent between an unlabeled example and an augmented unlabeled example. Unlike previous methods that use random noise such as Gaussian noise or dropout noise, UDA has a small twist in that it makes use of harder and more realistic noise generated by state-of-the-art data augmentation methods. This small twist leads to substantial improvements on six language tasks and three vision tasks even when the labeled set is extremely small. For example, on the IMDb text classification dataset, with only 20 labeled examples, UDA achieves an error rate of 4.20, outperforming the state-of-the-art model trained on 25,000 labeled examples. On standard semi-supervised learning benchmarks CIFAR-10 and SVHN, UDA outperforms all previous approaches and achieves an error rate of 2.7% on CIFAR-10 with only 4,000 examples and an error rate of 2.85% on SVHN with only 250 examples, nearly matching the performance of models trained on the full sets which are one or two orders of magnitude larger. UDA also works well on large-scale datasets such as ImageNet. When trained with 10% of the labeled set, UDA improves the top-1/top-5 accuracy from 55.1/77.3% to 68.7/88.5%. For the full ImageNet with 1.3M extra unlabeled data, UDA further pushes the performance from 78.3/94.4% to 79.0/94.5%.
Biomedical image segmentation is an important task in many medical applications. Segmentation methods based on convolutional neural networks attain state-of-the-art accuracy; however, they typically rely on supervised training with large labeled datasets. Labeling datasets of medical images requires significant expertise and time, and is infeasible at large scales. To tackle the lack of labeled data, researchers use techniques such as hand-engineered preprocessing steps, hand-tuned architectures, and data augmentation. However, these techniques involve costly engineering efforts, and are typically dataset-specific. We present an automated data augmentation method for medical images. We demonstrate our method on the task of segmenting magnetic resonance imaging (MRI) brain scans, focusing on the one-shot segmentation scenario -- a practical challenge in many medical applications. Our method requires only a single segmented scan, and leverages other unlabeled scans in a semi-supervised approach. We learn a model of transforms from the images, and use the model along with the labeled example to synthesize additional labeled training examples for supervised segmentation. Each transform is comprised of a spatial deformation field and an intensity change, enabling the synthesis of complex effects such as variations in anatomy and image acquisition procedures. Augmenting the training of a supervised segmenter with these new examples provides significant improvements over state-of-the-art methods for one-shot biomedical image segmentation. Our code is available at //github.com/xamyzhao/brainstorm.
Semantic segmentation is one of the basic topics in computer vision, it aims to assign semantic labels to every pixel of an image. Unbalanced semantic label distribution could have a negative influence on segmentation accuracy. In this paper, we investigate using data augmentation approach to balance the semantic label distribution in order to improve segmentation performance. We propose using generative adversarial networks (GANs) to generate realistic images for improving the performance of semantic segmentation networks. Experimental results show that the proposed method can not only improve segmentation performance on those classes with low accuracy, but also obtain 1.3% to 2.1% increase in average segmentation accuracy. It shows that this augmentation method can boost accuracy and be easily applicable to any other segmentation models.
Convolutional networks (ConvNets) have achieved great successes in various challenging vision tasks. However, the performance of ConvNets would degrade when encountering the domain shift. The domain adaptation is more significant while challenging in the field of biomedical image analysis, where cross-modality data have largely different distributions. Given that annotating the medical data is especially expensive, the supervised transfer learning approaches are not quite optimal. In this paper, we propose an unsupervised domain adaptation framework with adversarial learning for cross-modality biomedical image segmentations. Specifically, our model is based on a dilated fully convolutional network for pixel-wise prediction. Moreover, we build a plug-and-play domain adaptation module (DAM) to map the target input to features which are aligned with source domain feature space. A domain critic module (DCM) is set up for discriminating the feature space of both domains. We optimize the DAM and DCM via an adversarial loss without using any target domain label. Our proposed method is validated by adapting a ConvNet trained with MRI images to unpaired CT data for cardiac structures segmentations, and achieved very promising results.
Deep Convolutional Neural Networks have pushed the state-of-the art for semantic segmentation provided that a large amount of images together with pixel-wise annotations is available. Data collection is expensive and a solution to alleviate it is to use transfer learning. This reduces the amount of annotated data required for the network training but it does not get rid of this heavy processing step. We propose a method of transfer learning without annotations on the target task for datasets with redundant content and distinct pixel distributions. Our method takes advantage of the approximate content alignment of the images between two datasets when the approximation error prevents the reuse of annotation from one dataset to another. Given the annotations for only one dataset, we train a first network in a supervised manner. This network autonomously learns to generate deep data representations relevant to the semantic segmentation. Then the images in the new dataset, we train a new network to generate a deep data representation that matches the one from the first network on the previous dataset. The training consists in a regression between feature maps and does not require any annotations on the new dataset. We show that this method reaches performances similar to a classic transfer learning on the PASCAL VOC dataset with synthetic transformations.
We propose a novel locally adaptive learning estimator for enhancing the inter- and intra- discriminative capabilities of Deep Neural Networks, which can be used as improved loss layer for semantic image segmentation tasks. Most loss layers compute pixel-wise cost between feature maps and ground truths, ignoring spatial layouts and interactions between neighboring pixels with same object category, and thus networks cannot be effectively sensitive to intra-class connections. Stride by stride, our method firstly conducts adaptive pooling filter operating over predicted feature maps, aiming to merge predicted distributions over a small group of neighboring pixels with same category, and then it computes cost between the merged distribution vector and their category label. Such design can make groups of neighboring predictions from same category involved into estimations on predicting correctness with respect to their category, and hence train networks to be more sensitive to regional connections between adjacent pixels based on their categories. In the experiments on Pascal VOC 2012 segmentation datasets, the consistently improved results show that our proposed approach achieves better segmentation masks against previous counterparts.