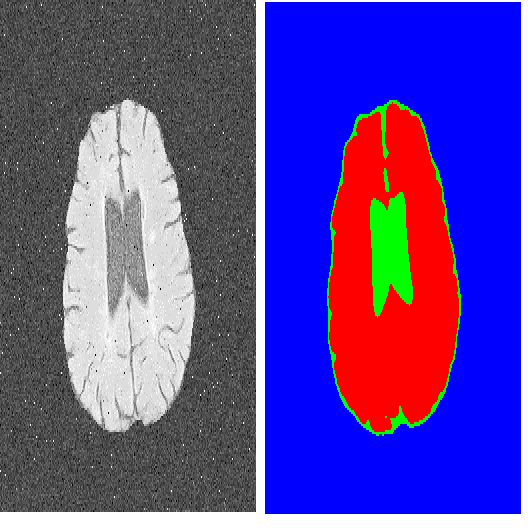
Transformers have made remarkable progress towards modeling long-range dependencies within the medical image analysis domain. However, current transformer-based models suffer from several disadvantages: 1) existing methods fail to capture the important features of the images due to the naive tokenization scheme; 2) the models suffer from information loss because they only consider single-scale feature representations; and 3) the segmentation label maps generated by the models are not accurate enough without considering rich semantic contexts and anatomical textures. In this work, we present CA-GANformer, a novel type of generative adversarial transformers, for medical image segmentation. First, we take advantage of the pyramid structure to construct multi-scale representations and handle multi-scale variations. We then design a novel class-aware transformer module to better learn the discriminative regions of objects with semantic structures. Lastly, we utilize an adversarial training strategy that boosts segmentation accuracy and correspondingly allows a transformer-based discriminator to capture high-level semantically correlated contents and low-level anatomical features. Our experiments demonstrate that CA-GANformer dramatically outperforms previous state-of-the-art transformer-based approaches on three benchmarks, obtaining absolute 2.54%-5.88% improvements in Dice over previous models. Further qualitative experiments provide a more detailed picture of the model's inner workings, shed light on the challenges in improved transparency, and demonstrate that transfer learning can greatly improve performance and reduce the size of medical image datasets in training, making CA-GANformer a strong starting point for downstream medical image analysis tasks. Codes and models will be available to the public.
相關內容
In this paper, we propose a novel mutual consistency network (MC-Net+) to effectively exploit the unlabeled data for semi-supervised medical image segmentation. The MC-Net+ model is motivated by the observation that deep models trained with limited annotations are prone to output highly uncertain and easily mis-classified predictions in ambiguous regions (e.g., adhesive edges or thin branches) for medical image segmentation. Leveraging these region-level challenging samples can make the semi-supervised segmentation model training more effective. Therefore, our proposed MC-Net+ model consists of two new designs. First, the model contains one shared encoder and multiple slightly different decoders (i.e., using different up-sampling strategies). The statistical discrepancy of multiple decoders' outputs is computed to denote the model's uncertainty, which indicates the unlabeled hard regions. Second, we apply a novel mutual consistency constraint between one decoder's probability output and other decoders' soft pseudo labels. In this way, we minimize the discrepancy of multiple outputs (i.e., the model uncertainty) during training and force the model to generate invariant results in such challenging regions, aiming at capturing more useful features. We compared the segmentation results of our MC-Net+ with five state-of-the-art semi-supervised approaches on three public medical datasets. Extension experiments with two common semi-supervised settings demonstrate the superior performance of our model over other existing methods, which sets a new state of the art for semi-supervised medical image segmentation.
Accurate segmentation of power lines in various aerial images is very important for UAV flight safety. The complex background and very thin structures of power lines, however, make it an inherently difficult task in computer vision. This paper presents PLGAN, a simple yet effective method based on generative adversarial networks, to segment power lines from aerial images with different backgrounds. Instead of directly using the adversarial networks to generate the segmentation, we take their certain decoding features and embed them into another semantic segmentation network by considering more context, geometry, and appearance information of power lines. We further exploit the appropriate form of the generated images for high-quality feature embedding and define a new loss function in the Hough-transform parameter space to enhance the segmentation of very thin power lines. Extensive experiments and comprehensive analysis demonstrate that our proposed PLGAN outperforms the prior state-of-the-art methods for semantic segmentation and line detection.
Transformers have dominated the field of natural language processing, and recently impacted the computer vision area. In the field of medical image analysis, Transformers have also been successfully applied to full-stack clinical applications, including image synthesis/reconstruction, registration, segmentation, detection, and diagnosis. Our paper presents both a position paper and a primer, promoting awareness and application of Transformers in the field of medical image analysis. Specifically, we first overview the core concepts of the attention mechanism built into Transformers and other basic components. Second, we give a new taxonomy of various Transformer architectures tailored for medical image applications and discuss their limitations. Within this review, we investigate key challenges revolving around the use of Transformers in different learning paradigms, improving the model efficiency, and their coupling with other techniques. We hope this review can give a comprehensive picture of Transformers to the readers in the field of medical image analysis.
Applying artificial intelligence techniques in medical imaging is one of the most promising areas in medicine. However, most of the recent success in this area highly relies on large amounts of carefully annotated data, whereas annotating medical images is a costly process. In this paper, we propose a novel method, called FocalMix, which, to the best of our knowledge, is the first to leverage recent advances in semi-supervised learning (SSL) for 3D medical image detection. We conducted extensive experiments on two widely used datasets for lung nodule detection, LUNA16 and NLST. Results show that our proposed SSL methods can achieve a substantial improvement of up to 17.3% over state-of-the-art supervised learning approaches with 400 unlabeled CT scans.
We consider the problem of referring image segmentation. Given an input image and a natural language expression, the goal is to segment the object referred by the language expression in the image. Existing works in this area treat the language expression and the input image separately in their representations. They do not sufficiently capture long-range correlations between these two modalities. In this paper, we propose a cross-modal self-attention (CMSA) module that effectively captures the long-range dependencies between linguistic and visual features. Our model can adaptively focus on informative words in the referring expression and important regions in the input image. In addition, we propose a gated multi-level fusion module to selectively integrate self-attentive cross-modal features corresponding to different levels in the image. This module controls the information flow of features at different levels. We validate the proposed approach on four evaluation datasets. Our proposed approach consistently outperforms existing state-of-the-art methods.
The U-Net was presented in 2015. With its straight-forward and successful architecture it quickly evolved to a commonly used benchmark in medical image segmentation. The adaptation of the U-Net to novel problems, however, comprises several degrees of freedom regarding the exact architecture, preprocessing, training and inference. These choices are not independent of each other and substantially impact the overall performance. The present paper introduces the nnU-Net ('no-new-Net'), which refers to a robust and self-adapting framework on the basis of 2D and 3D vanilla U-Nets. We argue the strong case for taking away superfluous bells and whistles of many proposed network designs and instead focus on the remaining aspects that make out the performance and generalizability of a method. We evaluate the nnU-Net in the context of the Medical Segmentation Decathlon challenge, which measures segmentation performance in ten disciplines comprising distinct entities, image modalities, image geometries and dataset sizes, with no manual adjustments between datasets allowed. At the time of manuscript submission, nnU-Net achieves the highest mean dice scores across all classes and seven phase 1 tasks (except class 1 in BrainTumour) in the online leaderboard of the challenge.
Deep neural network architectures have traditionally been designed and explored with human expertise in a long-lasting trial-and-error process. This process requires huge amount of time, expertise, and resources. To address this tedious problem, we propose a novel algorithm to optimally find hyperparameters of a deep network architecture automatically. We specifically focus on designing neural architectures for medical image segmentation task. Our proposed method is based on a policy gradient reinforcement learning for which the reward function is assigned a segmentation evaluation utility (i.e., dice index). We show the efficacy of the proposed method with its low computational cost in comparison with the state-of-the-art medical image segmentation networks. We also present a new architecture design, a densely connected encoder-decoder CNN, as a strong baseline architecture to apply the proposed hyperparameter search algorithm. We apply the proposed algorithm to each layer of the baseline architectures. As an application, we train the proposed system on cine cardiac MR images from Automated Cardiac Diagnosis Challenge (ACDC) MICCAI 2017. Starting from a baseline segmentation architecture, the resulting network architecture obtains the state-of-the-art results in accuracy without performing any trial-and-error based architecture design approaches or close supervision of the hyperparameters changes.
Convolutional networks (ConvNets) have achieved great successes in various challenging vision tasks. However, the performance of ConvNets would degrade when encountering the domain shift. The domain adaptation is more significant while challenging in the field of biomedical image analysis, where cross-modality data have largely different distributions. Given that annotating the medical data is especially expensive, the supervised transfer learning approaches are not quite optimal. In this paper, we propose an unsupervised domain adaptation framework with adversarial learning for cross-modality biomedical image segmentations. Specifically, our model is based on a dilated fully convolutional network for pixel-wise prediction. Moreover, we build a plug-and-play domain adaptation module (DAM) to map the target input to features which are aligned with source domain feature space. A domain critic module (DCM) is set up for discriminating the feature space of both domains. We optimize the DAM and DCM via an adversarial loss without using any target domain label. Our proposed method is validated by adapting a ConvNet trained with MRI images to unpaired CT data for cardiac structures segmentations, and achieved very promising results.

Recent advances in 3D fully convolutional networks (FCN) have made it feasible to produce dense voxel-wise predictions of volumetric images. In this work, we show that a multi-class 3D FCN trained on manually labeled CT scans of several anatomical structures (ranging from the large organs to thin vessels) can achieve competitive segmentation results, while avoiding the need for handcrafting features or training class-specific models. To this end, we propose a two-stage, coarse-to-fine approach that will first use a 3D FCN to roughly define a candidate region, which will then be used as input to a second 3D FCN. This reduces the number of voxels the second FCN has to classify to ~10% and allows it to focus on more detailed segmentation of the organs and vessels. We utilize training and validation sets consisting of 331 clinical CT images and test our models on a completely unseen data collection acquired at a different hospital that includes 150 CT scans, targeting three anatomical organs (liver, spleen, and pancreas). In challenging organs such as the pancreas, our cascaded approach improves the mean Dice score from 68.5 to 82.2%, achieving the highest reported average score on this dataset. We compare with a 2D FCN method on a separate dataset of 240 CT scans with 18 classes and achieve a significantly higher performance in small organs and vessels. Furthermore, we explore fine-tuning our models to different datasets. Our experiments illustrate the promise and robustness of current 3D FCN based semantic segmentation of medical images, achieving state-of-the-art results. Our code and trained models are available for download: //github.com/holgerroth/3Dunet_abdomen_cascade.
High spectral dimensionality and the shortage of annotations make hyperspectral image (HSI) classification a challenging problem. Recent studies suggest that convolutional neural networks can learn discriminative spatial features, which play a paramount role in HSI interpretation. However, most of these methods ignore the distinctive spectral-spatial characteristic of hyperspectral data. In addition, a large amount of unlabeled data remains an unexploited gold mine for efficient data use. Therefore, we proposed an integration of generative adversarial networks (GANs) and probabilistic graphical models for HSI classification. Specifically, we used a spectral-spatial generator and a discriminator to identify land cover categories of hyperspectral cubes. Moreover, to take advantage of a large amount of unlabeled data, we adopted a conditional random field to refine the preliminary classification results generated by GANs. Experimental results obtained using two commonly studied datasets demonstrate that the proposed framework achieved encouraging classification accuracy using a small number of data for training.