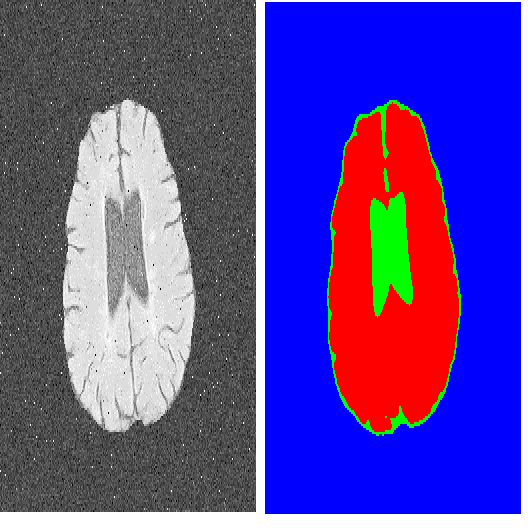
The new era of image segmentation leveraging the power of Deep Neural Nets (DNNs) comes with a price tag: to train a neural network for pixel-wise segmentation, a large amount of training samples has to be manually labeled on pixel-precision. In this work, we address this by following an indirect solution. We build upon the advances of the Explainable AI (XAI) community and extract a pixel-wise binary segmentation from the output of the Layer-wise Relevance Propagation (LRP) explaining the decision of a classification network. We show that we achieve similar results compared to an established U-Net segmentation architecture, while the generation of the training data is significantly simplified. The proposed method can be trained in a weakly supervised fashion, as the training samples must be only labeled on image-level, at the same time enabling the output of a segmentation mask. This makes it especially applicable to a wider range of real applications where tedious pixel-level labelling is often not possible.
相關內容
In this paper, we propose a novel mutual consistency network (MC-Net+) to effectively exploit the unlabeled data for semi-supervised medical image segmentation. The MC-Net+ model is motivated by the observation that deep models trained with limited annotations are prone to output highly uncertain and easily mis-classified predictions in ambiguous regions (e.g., adhesive edges or thin branches) for medical image segmentation. Leveraging these region-level challenging samples can make the semi-supervised segmentation model training more effective. Therefore, our proposed MC-Net+ model consists of two new designs. First, the model contains one shared encoder and multiple slightly different decoders (i.e., using different up-sampling strategies). The statistical discrepancy of multiple decoders' outputs is computed to denote the model's uncertainty, which indicates the unlabeled hard regions. Second, we apply a novel mutual consistency constraint between one decoder's probability output and other decoders' soft pseudo labels. In this way, we minimize the discrepancy of multiple outputs (i.e., the model uncertainty) during training and force the model to generate invariant results in such challenging regions, aiming at capturing more useful features. We compared the segmentation results of our MC-Net+ with five state-of-the-art semi-supervised approaches on three public medical datasets. Extension experiments with two common semi-supervised settings demonstrate the superior performance of our model over other existing methods, which sets a new state of the art for semi-supervised medical image segmentation.
We introduce a segmentation-guided approach to synthesise images that integrate features from two distinct domains. Images synthesised by our dual-domain model belong to one domain within the semantic mask, and to another in the rest of the image - smoothly integrated. We build on the successes of few-shot StyleGAN and single-shot semantic segmentation to minimise the amount of training required in utilising two domains. The method combines a few-shot cross-domain StyleGAN with a latent optimiser to achieve images containing features of two distinct domains. We use a segmentation-guided perceptual loss, which compares both pixel-level and activations between domain-specific and dual-domain synthetic images. Results demonstrate qualitatively and quantitatively that our model is capable of synthesising dual-domain images on a variety of objects (faces, horses, cats, cars), domains (natural, caricature, sketches) and part-based masks (eyes, nose, mouth, hair, car bonnet). The code is publicly available at: //github.com/denabazazian/Dual-Domain-Synthesis.
Deep learning has achieved remarkable results in many computer vision tasks. Deep neural networks typically rely on large amounts of training data to avoid overfitting. However, labeled data for real-world applications may be limited. By improving the quantity and diversity of training data, data augmentation has become an inevitable part of deep learning model training with image data. As an effective way to improve the sufficiency and diversity of training data, data augmentation has become a necessary part of successful application of deep learning models on image data. In this paper, we systematically review different image data augmentation methods. We propose a taxonomy of reviewed methods and present the strengths and limitations of these methods. We also conduct extensive experiments with various data augmentation methods on three typical computer vision tasks, including semantic segmentation, image classification and object detection. Finally, we discuss current challenges faced by data augmentation and future research directions to put forward some useful research guidance.
Image segmentation is a key topic in image processing and computer vision with applications such as scene understanding, medical image analysis, robotic perception, video surveillance, augmented reality, and image compression, among many others. Various algorithms for image segmentation have been developed in the literature. Recently, due to the success of deep learning models in a wide range of vision applications, there has been a substantial amount of works aimed at developing image segmentation approaches using deep learning models. In this survey, we provide a comprehensive review of the literature at the time of this writing, covering a broad spectrum of pioneering works for semantic and instance-level segmentation, including fully convolutional pixel-labeling networks, encoder-decoder architectures, multi-scale and pyramid based approaches, recurrent networks, visual attention models, and generative models in adversarial settings. We investigate the similarity, strengths and challenges of these deep learning models, examine the most widely used datasets, report performances, and discuss promising future research directions in this area.
A key requirement for the success of supervised deep learning is a large labeled dataset - a condition that is difficult to meet in medical image analysis. Self-supervised learning (SSL) can help in this regard by providing a strategy to pre-train a neural network with unlabeled data, followed by fine-tuning for a downstream task with limited annotations. Contrastive learning, a particular variant of SSL, is a powerful technique for learning image-level representations. In this work, we propose strategies for extending the contrastive learning framework for segmentation of volumetric medical images in the semi-supervised setting with limited annotations, by leveraging domain-specific and problem-specific cues. Specifically, we propose (1) novel contrasting strategies that leverage structural similarity across volumetric medical images (domain-specific cue) and (2) a local version of the contrastive loss to learn distinctive representations of local regions that are useful for per-pixel segmentation (problem-specific cue). We carry out an extensive evaluation on three Magnetic Resonance Imaging (MRI) datasets. In the limited annotation setting, the proposed method yields substantial improvements compared to other self-supervision and semi-supervised learning techniques. When combined with a simple data augmentation technique, the proposed method reaches within 8% of benchmark performance using only two labeled MRI volumes for training, corresponding to only 4% (for ACDC) of the training data used to train the benchmark.
Applying artificial intelligence techniques in medical imaging is one of the most promising areas in medicine. However, most of the recent success in this area highly relies on large amounts of carefully annotated data, whereas annotating medical images is a costly process. In this paper, we propose a novel method, called FocalMix, which, to the best of our knowledge, is the first to leverage recent advances in semi-supervised learning (SSL) for 3D medical image detection. We conducted extensive experiments on two widely used datasets for lung nodule detection, LUNA16 and NLST. Results show that our proposed SSL methods can achieve a substantial improvement of up to 17.3% over state-of-the-art supervised learning approaches with 400 unlabeled CT scans.
Few-shot image classification aims to classify unseen classes with limited labeled samples. Recent works benefit from the meta-learning process with episodic tasks and can fast adapt to class from training to testing. Due to the limited number of samples for each task, the initial embedding network for meta learning becomes an essential component and can largely affects the performance in practice. To this end, many pre-trained methods have been proposed, and most of them are trained in supervised way with limited transfer ability for unseen classes. In this paper, we proposed to train a more generalized embedding network with self-supervised learning (SSL) which can provide slow and robust representation for downstream tasks by learning from the data itself. We evaluate our work by extensive comparisons with previous baseline methods on two few-shot classification datasets ({\em i.e.,} MiniImageNet and CUB). Based on the evaluation results, the proposed method achieves significantly better performance, i.e., improve 1-shot and 5-shot tasks by nearly \textbf{3\%} and \textbf{4\%} on MiniImageNet, by nearly \textbf{9\%} and \textbf{3\%} on CUB. Moreover, the proposed method can gain the improvement of (\textbf{15\%}, \textbf{13\%}) on MiniImageNet and (\textbf{15\%}, \textbf{8\%}) on CUB by pretraining using more unlabeled data. Our code will be available at \hyperref[//github.com/phecy/SSL-FEW-SHOT.]{//github.com/phecy/ssl-few-shot.}
The U-Net was presented in 2015. With its straight-forward and successful architecture it quickly evolved to a commonly used benchmark in medical image segmentation. The adaptation of the U-Net to novel problems, however, comprises several degrees of freedom regarding the exact architecture, preprocessing, training and inference. These choices are not independent of each other and substantially impact the overall performance. The present paper introduces the nnU-Net ('no-new-Net'), which refers to a robust and self-adapting framework on the basis of 2D and 3D vanilla U-Nets. We argue the strong case for taking away superfluous bells and whistles of many proposed network designs and instead focus on the remaining aspects that make out the performance and generalizability of a method. We evaluate the nnU-Net in the context of the Medical Segmentation Decathlon challenge, which measures segmentation performance in ten disciplines comprising distinct entities, image modalities, image geometries and dataset sizes, with no manual adjustments between datasets allowed. At the time of manuscript submission, nnU-Net achieves the highest mean dice scores across all classes and seven phase 1 tasks (except class 1 in BrainTumour) in the online leaderboard of the challenge.
Deep Convolutional Neural Networks have pushed the state-of-the art for semantic segmentation provided that a large amount of images together with pixel-wise annotations is available. Data collection is expensive and a solution to alleviate it is to use transfer learning. This reduces the amount of annotated data required for the network training but it does not get rid of this heavy processing step. We propose a method of transfer learning without annotations on the target task for datasets with redundant content and distinct pixel distributions. Our method takes advantage of the approximate content alignment of the images between two datasets when the approximation error prevents the reuse of annotation from one dataset to another. Given the annotations for only one dataset, we train a first network in a supervised manner. This network autonomously learns to generate deep data representations relevant to the semantic segmentation. Then the images in the new dataset, we train a new network to generate a deep data representation that matches the one from the first network on the previous dataset. The training consists in a regression between feature maps and does not require any annotations on the new dataset. We show that this method reaches performances similar to a classic transfer learning on the PASCAL VOC dataset with synthetic transformations.

Recent advances in 3D fully convolutional networks (FCN) have made it feasible to produce dense voxel-wise predictions of volumetric images. In this work, we show that a multi-class 3D FCN trained on manually labeled CT scans of several anatomical structures (ranging from the large organs to thin vessels) can achieve competitive segmentation results, while avoiding the need for handcrafting features or training class-specific models. To this end, we propose a two-stage, coarse-to-fine approach that will first use a 3D FCN to roughly define a candidate region, which will then be used as input to a second 3D FCN. This reduces the number of voxels the second FCN has to classify to ~10% and allows it to focus on more detailed segmentation of the organs and vessels. We utilize training and validation sets consisting of 331 clinical CT images and test our models on a completely unseen data collection acquired at a different hospital that includes 150 CT scans, targeting three anatomical organs (liver, spleen, and pancreas). In challenging organs such as the pancreas, our cascaded approach improves the mean Dice score from 68.5 to 82.2%, achieving the highest reported average score on this dataset. We compare with a 2D FCN method on a separate dataset of 240 CT scans with 18 classes and achieve a significantly higher performance in small organs and vessels. Furthermore, we explore fine-tuning our models to different datasets. Our experiments illustrate the promise and robustness of current 3D FCN based semantic segmentation of medical images, achieving state-of-the-art results. Our code and trained models are available for download: //github.com/holgerroth/3Dunet_abdomen_cascade.