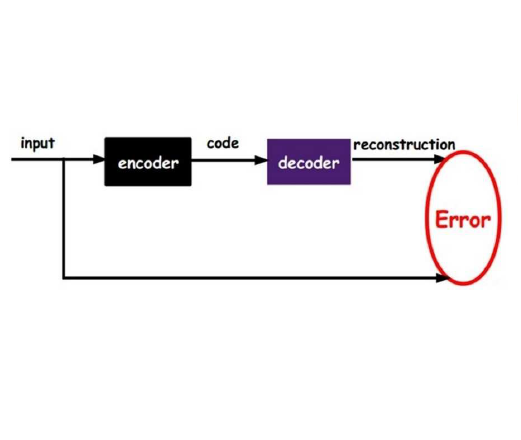
Despite the increasing use of deep learning in medical image segmentation, acquiring sufficient training data remains a challenge in the medical field. In response, data augmentation techniques have been proposed; however, the generation of diverse and realistic medical images and their corresponding masks remains a difficult task, especially when working with insufficient training sets. To address these limitations, we present an end-to-end architecture based on the Hamiltonian Variational Autoencoder (HVAE). This approach yields an improved posterior distribution approximation compared to traditional Variational Autoencoders (VAE), resulting in higher image generation quality. Our method outperforms generative adversarial architectures under data-scarce conditions, showcasing enhancements in image quality and precise tumor mask synthesis. We conduct experiments on two publicly available datasets, MICCAI's Brain Tumor Segmentation Challenge (BRATS), and Head and Neck Tumor Segmentation Challenge (HECKTOR), demonstrating the effectiveness of our method on different medical imaging modalities.
相關內容
To handle the large scale of whole slide images in computational pathology, most approaches first tessellate the images into smaller patches, extract features from these patches, and finally aggregate the feature vectors with weakly-supervised learning. The performance of this workflow strongly depends on the quality of the extracted features. Recently, foundation models in computer vision showed that leveraging huge amounts of data through supervised or self-supervised learning improves feature quality and generalizability for a variety of tasks. In this study, we benchmark the most popular vision foundation models as feature extractors for histopathology data. We evaluate the models in two settings: slide-level classification and patch-level classification. We show that foundation models are a strong baseline. Our experiments demonstrate that by finetuning a foundation model on a single GPU for only two hours or three days depending on the dataset, we can match or outperform state-of-the-art feature extractors for computational pathology. These findings imply that even with little resources one can finetune a feature extractor tailored towards a specific downstream task and dataset. This is a considerable shift from the current state, where only few institutions with large amounts of resources and datasets are able to train a feature extractor. We publish all code used for training and evaluation as well as the finetuned models.
Modelling of systems where the full system information is unknown is an oft encountered problem for various engineering and industrial applications, as it's either impossible to consider all the complex physics involved or simpler models are considered to keep within the limits of the available resources. Recent advances in greybox modelling like the deep hidden physics models address this space by combining data and physics. However, for most real-life applications, model generalizability is a key issue, as retraining a model for every small change in system inputs and parameters or modification in domain configuration can render the model economically unviable. In this work we present a novel enhancement to the idea of hidden physics models which can generalize for changes in system inputs, parameters and domains. We also show that this approach holds promise in system discovery as well and helps learn the hidden physics for the changed system inputs, parameters and domain configuration.
Utilizing massive web-scale datasets has led to unprecedented performance gains in machine learning models, but also imposes outlandish compute requirements for their training. In order to improve training and data efficiency, we here push the limits of pruning large-scale multimodal datasets for training CLIP-style models. Today's most effective pruning method on ImageNet clusters data samples into separate concepts according to their embedding and prunes away the most prototypical samples. We scale this approach to LAION and improve it by noting that the pruning rate should be concept-specific and adapted to the complexity of the concept. Using a simple and intuitive complexity measure, we are able to reduce the training cost to a quarter of regular training. By filtering from the LAION dataset, we find that training on a smaller set of high-quality data can lead to higher performance with significantly lower training costs. More specifically, we are able to outperform the LAION-trained OpenCLIP-ViT-B32 model on ImageNet zero-shot accuracy by 1.1p.p. while only using 27.7% of the data and training compute. Despite a strong reduction in training cost, we also see improvements on ImageNet dist. shifts, retrieval tasks and VTAB. On the DataComp Medium benchmark, we achieve a new state-of-the-art ImageNet zero-shot accuracy and a competitive average zero-shot accuracy on 38 evaluation tasks.
We present a system for anomaly detection in histopathological images. In histology, normal samples are usually abundant, whereas anomalous (pathological) cases are scarce or not available. Under such settings, one-class classifiers trained on healthy data can detect out-of-distribution anomalous samples. Such approaches combined with pre-trained Convolutional Neural Network (CNN) representations of images were previously employed for anomaly detection (AD). However, pre-trained off-the-shelf CNN representations may not be sensitive to abnormal conditions in tissues, while natural variations of healthy tissue may result in distant representations. To adapt representations to relevant details in healthy tissue we propose training a CNN on an auxiliary task that discriminates healthy tissue of different species, organs, and staining reagents. Almost no additional labeling workload is required, since healthy samples come automatically with aforementioned labels. During training we enforce compact image representations with a center-loss term, which further improves representations for AD. The proposed system outperforms established AD methods on a published dataset of liver anomalies. Moreover, it provided comparable results to conventional methods specifically tailored for quantification of liver anomalies. We show that our approach can be used for toxicity assessment of candidate drugs at early development stages and thereby may reduce expensive late-stage drug attrition.
In many medical image classification problems, labeled data is scarce while unlabeled data is more available. Semi-supervised learning and self-supervised learning are two different research directions that can improve accuracy by learning from extra unlabeled data. Recent methods from both directions have reported significant gains on traditional benchmarks. Yet past benchmarks do not focus on medical tasks and rarely compare self- and semi- methods together on equal footing. Furthermore, past benchmarks often handle hyperparameter tuning suboptimally. First, they may not tune hyperparameters at all, leading to underfitting. Second, when tuning does occur, it often unrealistically uses a labeled validation set much larger than the train set. Both cases make previously published rankings of methods difficult to translate to practical settings. This study contributes a systematic evaluation of self- and semi- methods with a unified experimental protocol intended to guide a practitioner with scarce overall labeled data and a limited compute budget. We answer two key questions: Can hyperparameter tuning be effective with realistic-sized validation sets? If so, when all methods are tuned well, which self- or semi-supervised methods reach the best accuracy? Our study compares 13 representative semi- and self-supervised methods to strong labeled-set-only baselines on 4 medical datasets. From 20000+ total GPU hours of computation, we provide valuable best practices to resource-constrained, results-focused practitioners.
In designing external validation studies of clinical prediction models, contemporary sample size calculation methods are based on the frequentist inferential paradigm. One of the widely reported metrics of model performance is net benefit (NB), and the relevance of conventional inference around NB as a measure of clinical utility is doubtful. Value of Information methodology quantifies the consequences of uncertainty in terms of its impact on clinical utility of decisions. We introduce the expected value of sample information (EVSI) for validation as the expected gain in NB from conducting an external validation study of a given size. We propose algorithms for EVSI computation, and in a case study demonstrate how EVSI changes as a function of the amount of current information and future study's sample size. Value of Information methodology provides a decision-theoretic lens to the process of planning a validation study of a risk prediction model and can complement conventional methods when designing such studies.
Large annotated datasets are required for training deep learning models, but in medical imaging data sharing is often complicated due to ethics, anonymization and data protection legislation. Generative AI models, such as generative adversarial networks (GANs) and diffusion models, can today produce very realistic synthetic images, and can potentially facilitate data sharing. However, in order to share synthetic medical images it must first be demonstrated that they can be used for training different networks with acceptable performance. Here, we therefore comprehensively evaluate four GANs (progressive GAN, StyleGAN 1-3) and a diffusion model for the task of brain tumor segmentation (using two segmentation networks, U-Net and a Swin transformer). Our results show that segmentation networks trained on synthetic images reach Dice scores that are 80% - 90% of Dice scores when training with real images, but that memorization of the training images can be a problem for diffusion models if the original dataset is too small. Our conclusion is that sharing synthetic medical images is a viable option to sharing real images, but that further work is required. The trained generative models and the generated synthetic images are shared on AIDA data hub
We hypothesize that due to the greedy nature of learning in multi-modal deep neural networks, these models tend to rely on just one modality while under-fitting the other modalities. Such behavior is counter-intuitive and hurts the models' generalization, as we observe empirically. To estimate the model's dependence on each modality, we compute the gain on the accuracy when the model has access to it in addition to another modality. We refer to this gain as the conditional utilization rate. In the experiments, we consistently observe an imbalance in conditional utilization rates between modalities, across multiple tasks and architectures. Since conditional utilization rate cannot be computed efficiently during training, we introduce a proxy for it based on the pace at which the model learns from each modality, which we refer to as the conditional learning speed. We propose an algorithm to balance the conditional learning speeds between modalities during training and demonstrate that it indeed addresses the issue of greedy learning. The proposed algorithm improves the model's generalization on three datasets: Colored MNIST, Princeton ModelNet40, and NVIDIA Dynamic Hand Gesture.
A key requirement for the success of supervised deep learning is a large labeled dataset - a condition that is difficult to meet in medical image analysis. Self-supervised learning (SSL) can help in this regard by providing a strategy to pre-train a neural network with unlabeled data, followed by fine-tuning for a downstream task with limited annotations. Contrastive learning, a particular variant of SSL, is a powerful technique for learning image-level representations. In this work, we propose strategies for extending the contrastive learning framework for segmentation of volumetric medical images in the semi-supervised setting with limited annotations, by leveraging domain-specific and problem-specific cues. Specifically, we propose (1) novel contrasting strategies that leverage structural similarity across volumetric medical images (domain-specific cue) and (2) a local version of the contrastive loss to learn distinctive representations of local regions that are useful for per-pixel segmentation (problem-specific cue). We carry out an extensive evaluation on three Magnetic Resonance Imaging (MRI) datasets. In the limited annotation setting, the proposed method yields substantial improvements compared to other self-supervision and semi-supervised learning techniques. When combined with a simple data augmentation technique, the proposed method reaches within 8% of benchmark performance using only two labeled MRI volumes for training, corresponding to only 4% (for ACDC) of the training data used to train the benchmark.
Graph representation learning for hypergraphs can be used to extract patterns among higher-order interactions that are critically important in many real world problems. Current approaches designed for hypergraphs, however, are unable to handle different types of hypergraphs and are typically not generic for various learning tasks. Indeed, models that can predict variable-sized heterogeneous hyperedges have not been available. Here we develop a new self-attention based graph neural network called Hyper-SAGNN applicable to homogeneous and heterogeneous hypergraphs with variable hyperedge sizes. We perform extensive evaluations on multiple datasets, including four benchmark network datasets and two single-cell Hi-C datasets in genomics. We demonstrate that Hyper-SAGNN significantly outperforms the state-of-the-art methods on traditional tasks while also achieving great performance on a new task called outsider identification. Hyper-SAGNN will be useful for graph representation learning to uncover complex higher-order interactions in different applications.