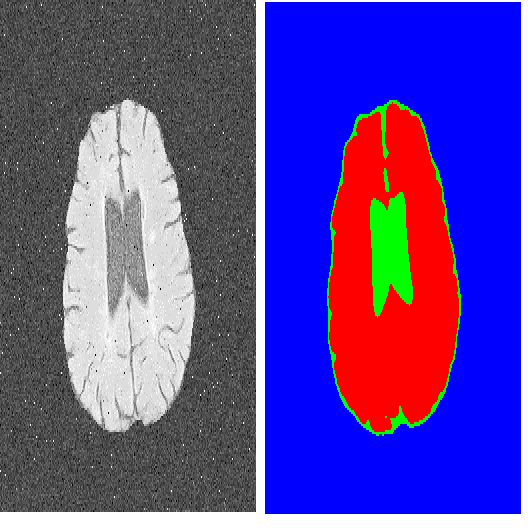
The Segment Anything Model (SAM) is the first foundation model for general image segmentation. It designed a novel promotable segmentation task, ensuring zero-shot image segmentation using the pre-trained model via two main modes including automatic everything and manual prompt. SAM has achieved impressive results on various natural image segmentation tasks. However, medical image segmentation (MIS) is more challenging due to the complex modalities, fine anatomical structures, uncertain and complex object boundaries, and wide-range object scales. SAM has achieved impressive results on various natural image segmentation tasks. Meanwhile, zero-shot and efficient MIS can well reduce the annotation time and boost the development of medical image analysis. Hence, SAM seems to be a potential tool and its performance on large medical datasets should be further validated. We collected and sorted 52 open-source datasets, and build a large medical segmentation dataset with 16 modalities, 68 objects, and 553K slices. We conducted a comprehensive analysis of different SAM testing strategies on the so-called COSMOS 553K dataset. Extensive experiments validate that SAM performs better with manual hints like points and boxes for object perception in medical images, leading to better performance in prompt mode compared to everything mode. Additionally, SAM shows remarkable performance in some specific objects and modalities, but is imperfect or even totally fails in other situations. Finally, we analyze the influence of different factors (e.g., the Fourier-based boundary complexity and size of the segmented objects) on SAM's segmentation performance. Extensive experiments validate that SAM's zero-shot segmentation capability is not sufficient to ensure its direct application to the MIS.
相關內容
Unsupervised object-centric representation (OCR) learning has recently drawn attention as a new paradigm of visual representation. This is because of its potential of being an effective pre-training technique for various downstream tasks in terms of sample efficiency, systematic generalization, and reasoning. Although image-based reinforcement learning (RL) is one of the most important and thus frequently mentioned such downstream tasks, the benefit in RL has surprisingly not been investigated systematically thus far. Instead, most of the evaluations have focused on rather indirect metrics such as segmentation quality and object property prediction accuracy. In this paper, we investigate the effectiveness of OCR pre-training for image-based reinforcement learning via empirical experiments. For systematic evaluation, we introduce a simple object-centric visual RL benchmark and conduct experiments to answer questions such as ``Does OCR pre-training improve performance on object-centric tasks?'' and ``Can OCR pre-training help with out-of-distribution generalization?''. Our results provide empirical evidence for valuable insights into the effectiveness of OCR pre-training for RL and the potential limitations of its use in certain scenarios. Additionally, this study also examines the critical aspects of incorporating OCR pre-training in RL, including performance in a visually complex environment and the appropriate pooling layer to aggregate the object representations.
Attention-based vision models, such as Vision Transformer (ViT) and its variants, have shown promising performance in various computer vision tasks. However, these emerging architectures suffer from large model sizes and high computational costs, calling for efficient model compression solutions. To date, pruning ViTs has been well studied, while other compression strategies that have been widely applied in CNN compression, e.g., model factorization, is little explored in the context of ViT compression. This paper explores an efficient method for compressing vision transformers to enrich the toolset for obtaining compact attention-based vision models. Based on the new insight on the multi-head attention layer, we develop a highly efficient ViT compression solution, which outperforms the state-of-the-art pruning methods. For compressing DeiT-small and DeiT-base models on ImageNet, our proposed approach can achieve 0.45% and 0.76% higher top-1 accuracy even with fewer parameters. Our finding can also be applied to improve the customization efficiency of text-to-image diffusion models, with much faster training (up to $2.6\times$ speedup) and lower extra storage cost (up to $1927.5\times$ reduction) than the existing works.
We present the Recognize Anything Model (RAM): a strong foundation model for image tagging. RAM makes a substantial step for large models in computer vision, demonstrating the zero-shot ability to recognize any common category with high accuracy. RAM introduces a new paradigm for image tagging, leveraging large-scale image-text pairs for training instead of manual annotations. The development of RAM comprises four key steps. Firstly, annotation-free image tags are obtained at scale through automatic text semantic parsing. Subsequently, a preliminary model is trained for automatic annotation by unifying the caption and tagging tasks, supervised by the original texts and parsed tags, respectively. Thirdly, a data engine is employed to generate additional annotations and clean incorrect ones. Lastly, the model is retrained with the processed data and fine-tuned using a smaller but higher-quality dataset. We evaluate the tagging capabilities of RAM on numerous benchmarks and observe impressive zero-shot performance, significantly outperforming CLIP and BLIP. Remarkably, RAM even surpasses the fully supervised manners and exhibits competitive performance with the Google tagging API. We are releasing the RAM at \url{//recognize-anything.github.io/} to foster the advancements of large models in computer vision.
Medical image segmentation is a fundamental and critical step in many image-guided clinical approaches. Recent success of deep learning-based segmentation methods usually relies on a large amount of labeled data, which is particularly difficult and costly to obtain especially in the medical imaging domain where only experts can provide reliable and accurate annotations. Semi-supervised learning has emerged as an appealing strategy and been widely applied to medical image segmentation tasks to train deep models with limited annotations. In this paper, we present a comprehensive review of recently proposed semi-supervised learning methods for medical image segmentation and summarized both the technical novelties and empirical results. Furthermore, we analyze and discuss the limitations and several unsolved problems of existing approaches. We hope this review could inspire the research community to explore solutions for this challenge and further promote the developments in medical image segmentation field.
Artificial neural networks thrive in solving the classification problem for a particular rigid task, acquiring knowledge through generalized learning behaviour from a distinct training phase. The resulting network resembles a static entity of knowledge, with endeavours to extend this knowledge without targeting the original task resulting in a catastrophic forgetting. Continual learning shifts this paradigm towards networks that can continually accumulate knowledge over different tasks without the need to retrain from scratch. We focus on task incremental classification, where tasks arrive sequentially and are delineated by clear boundaries. Our main contributions concern 1) a taxonomy and extensive overview of the state-of-the-art, 2) a novel framework to continually determine the stability-plasticity trade-off of the continual learner, 3) a comprehensive experimental comparison of 11 state-of-the-art continual learning methods and 4 baselines. We empirically scrutinize method strengths and weaknesses on three benchmarks, considering Tiny Imagenet and large-scale unbalanced iNaturalist and a sequence of recognition datasets. We study the influence of model capacity, weight decay and dropout regularization, and the order in which the tasks are presented, and qualitatively compare methods in terms of required memory, computation time, and storage.
A key requirement for the success of supervised deep learning is a large labeled dataset - a condition that is difficult to meet in medical image analysis. Self-supervised learning (SSL) can help in this regard by providing a strategy to pre-train a neural network with unlabeled data, followed by fine-tuning for a downstream task with limited annotations. Contrastive learning, a particular variant of SSL, is a powerful technique for learning image-level representations. In this work, we propose strategies for extending the contrastive learning framework for segmentation of volumetric medical images in the semi-supervised setting with limited annotations, by leveraging domain-specific and problem-specific cues. Specifically, we propose (1) novel contrasting strategies that leverage structural similarity across volumetric medical images (domain-specific cue) and (2) a local version of the contrastive loss to learn distinctive representations of local regions that are useful for per-pixel segmentation (problem-specific cue). We carry out an extensive evaluation on three Magnetic Resonance Imaging (MRI) datasets. In the limited annotation setting, the proposed method yields substantial improvements compared to other self-supervision and semi-supervised learning techniques. When combined with a simple data augmentation technique, the proposed method reaches within 8% of benchmark performance using only two labeled MRI volumes for training, corresponding to only 4% (for ACDC) of the training data used to train the benchmark.
The rapid advancements in machine learning, graphics processing technologies and availability of medical imaging data has led to a rapid increase in use of machine learning models in the medical domain. This was exacerbated by the rapid advancements in convolutional neural network (CNN) based architectures, which were adopted by the medical imaging community to assist clinicians in disease diagnosis. Since the grand success of AlexNet in 2012, CNNs have been increasingly used in medical image analysis to improve the efficiency of human clinicians. In recent years, three-dimensional (3D) CNNs have been employed for analysis of medical images. In this paper, we trace the history of how the 3D CNN was developed from its machine learning roots, brief mathematical description of 3D CNN and the preprocessing steps required for medical images before feeding them to 3D CNNs. We review the significant research in the field of 3D medical imaging analysis using 3D CNNs (and its variants) in different medical areas such as classification, segmentation, detection, and localization. We conclude by discussing the challenges associated with the use of 3D CNNs in the medical imaging domain (and the use of deep learning models, in general) and possible future trends in the field.
The U-Net was presented in 2015. With its straight-forward and successful architecture it quickly evolved to a commonly used benchmark in medical image segmentation. The adaptation of the U-Net to novel problems, however, comprises several degrees of freedom regarding the exact architecture, preprocessing, training and inference. These choices are not independent of each other and substantially impact the overall performance. The present paper introduces the nnU-Net ('no-new-Net'), which refers to a robust and self-adapting framework on the basis of 2D and 3D vanilla U-Nets. We argue the strong case for taking away superfluous bells and whistles of many proposed network designs and instead focus on the remaining aspects that make out the performance and generalizability of a method. We evaluate the nnU-Net in the context of the Medical Segmentation Decathlon challenge, which measures segmentation performance in ten disciplines comprising distinct entities, image modalities, image geometries and dataset sizes, with no manual adjustments between datasets allowed. At the time of manuscript submission, nnU-Net achieves the highest mean dice scores across all classes and seven phase 1 tasks (except class 1 in BrainTumour) in the online leaderboard of the challenge.
Medical image segmentation requires consensus ground truth segmentations to be derived from multiple expert annotations. A novel approach is proposed that obtains consensus segmentations from experts using graph cuts (GC) and semi supervised learning (SSL). Popular approaches use iterative Expectation Maximization (EM) to estimate the final annotation and quantify annotator's performance. Such techniques pose the risk of getting trapped in local minima. We propose a self consistency (SC) score to quantify annotator consistency using low level image features. SSL is used to predict missing annotations by considering global features and local image consistency. The SC score also serves as the penalty cost in a second order Markov random field (MRF) cost function optimized using graph cuts to derive the final consensus label. Graph cut obtains a global maximum without an iterative procedure. Experimental results on synthetic images, real data of Crohn's disease patients and retinal images show our final segmentation to be accurate and more consistent than competing methods.
Deep learning (DL) based semantic segmentation methods have been providing state-of-the-art performance in the last few years. More specifically, these techniques have been successfully applied to medical image classification, segmentation, and detection tasks. One deep learning technique, U-Net, has become one of the most popular for these applications. In this paper, we propose a Recurrent Convolutional Neural Network (RCNN) based on U-Net as well as a Recurrent Residual Convolutional Neural Network (RRCNN) based on U-Net models, which are named RU-Net and R2U-Net respectively. The proposed models utilize the power of U-Net, Residual Network, as well as RCNN. There are several advantages of these proposed architectures for segmentation tasks. First, a residual unit helps when training deep architecture. Second, feature accumulation with recurrent residual convolutional layers ensures better feature representation for segmentation tasks. Third, it allows us to design better U-Net architecture with same number of network parameters with better performance for medical image segmentation. The proposed models are tested on three benchmark datasets such as blood vessel segmentation in retina images, skin cancer segmentation, and lung lesion segmentation. The experimental results show superior performance on segmentation tasks compared to equivalent models including U-Net and residual U-Net (ResU-Net).