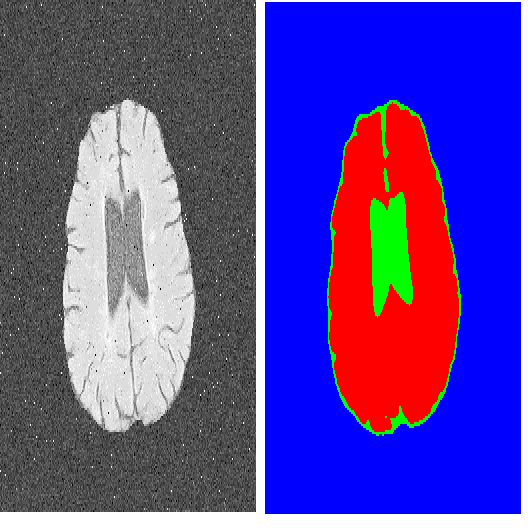
There has been exploding interest in embracing Transformer-based architectures for medical image segmentation. However, the lack of large-scale annotated medical datasets make achieving performances equivalent to those in natural images challenging. Convolutional networks, in contrast, have higher inductive biases and consequently, are easily trainable to high performance. Recently, the ConvNeXt architecture attempted to modernize the standard ConvNet by mirroring Transformer blocks. In this work, we improve upon this to design a modernized and scalable convolutional architecture customized to challenges of data-scarce medical settings. We introduce MedNeXt, a Transformer-inspired large kernel segmentation network which introduces - 1) A fully ConvNeXt 3D Encoder-Decoder Network for medical image segmentation, 2) Residual ConvNeXt up and downsampling blocks to preserve semantic richness across scales, 3) A novel technique to iteratively increase kernel sizes by upsampling small kernel networks, to prevent performance saturation on limited medical data, 4) Compound scaling at multiple levels (depth, width, kernel size) of MedNeXt. This leads to state-of-the-art performance on 4 tasks on CT and MRI modalities and varying dataset sizes, representing a modernized deep architecture for medical image segmentation.
相關內容
Over the past few years, the rapid development of deep learning technologies for computer vision has significantly improved the performance of medical image segmentation (MedISeg). However, the diverse implementation strategies of various models have led to an extremely complex MedISeg system, resulting in a potential problem of unfair result comparisons. In this paper, we collect a series of MedISeg tricks for different model implementation phases (i.e., pre-training model, data pre-processing, data augmentation, model implementation, model inference, and result post-processing), and experimentally explore the effectiveness of these tricks on consistent baselines. With the extensive experimental results on both the representative 2D and 3D medical image datasets, we explicitly clarify the effect of these tricks. Moreover, based on the surveyed tricks, we also open-sourced a strong MedISeg repository, where each component has the advantage of plug-and-play. We believe that this milestone work not only completes a comprehensive and complementary survey of the state-of-the-art MedISeg approaches, but also offers a practical guide for addressing the future medical image processing challenges including but not limited to small dataset, class imbalance learning, multi-modality learning, and domain adaptation. The code and training weights have been released at: //github.com/hust-linyi/seg_trick.
Achieving high-quality semantic segmentation predictions using only image-level labels enables a new level of real-world applicability. Although state-of-the-art networks deliver reliable predictions, the amount of handcrafted pixel-wise annotations to enable these results are not feasible in many real-world applications. Hence, several works have already targeted this bottleneck, using classifier-based networks like Class Activation Maps~\cite{CAM} (CAMs) as a base. Addressing CAM's weaknesses of fuzzy borders and incomplete predictions, state-of-the-art approaches rely only on adding regulations to the classifier loss or using pixel-similarity-based refinement after the fact. We propose a framework that introduces an additional module using object perimeters for improved saliency. We define object perimeter information as the line separating the object and background. Our new PerimeterFit module will be applied to pre-refine the CAM predictions before using the pixel-similarity-based network. In this way, our PerimeterFit increases the quality of the CAM prediction while simultaneously improving the false negative rate. We investigated a wide range of state-of-the-art unsupervised semantic segmentation networks and edge detection techniques to create useful perimeter maps, which enable our framework to predict object locations with sharper perimeters. We achieved up to 1.5% improvement over frameworks without our PerimeterFit module. We conduct an exhaustive analysis to illustrate that SILOP enhances existing state-of-the-art frameworks for image-level-based semantic segmentation. The framework is open-source and accessible online at //github.com/ErikOstrowski/SILOP.
Semantic segmentation has a broad range of applications in a variety of domains including land coverage analysis, autonomous driving, and medical image analysis. Convolutional neural networks (CNN) and Vision Transformers (ViTs) provide the architecture models for semantic segmentation. Even though ViTs have proven success in image classification, they cannot be directly applied to dense prediction tasks such as image segmentation and object detection since ViT is not a general purpose backbone due to its patch partitioning scheme. In this survey, we discuss some of the different ViT architectures that can be used for semantic segmentation and how their evolution managed the above-stated challenge. The rise of ViT and its performance with a high success rate motivated the community to slowly replace the traditional convolutional neural networks in various computer vision tasks. This survey aims to review and compare the performances of ViT architectures designed for semantic segmentation using benchmarking datasets. This will be worthwhile for the community to yield knowledge regarding the implementations carried out in semantic segmentation and to discover more efficient methodologies using ViTs.
The rapid advances in Vision Transformer (ViT) refresh the state-of-the-art performances in various vision tasks, overshadowing the conventional CNN-based models. This ignites a few recent striking-back research in the CNN world showing that pure CNN models can achieve as good performance as ViT models when carefully tuned. While encouraging, designing such high-performance CNN models is challenging, requiring non-trivial prior knowledge of network design. To this end, a novel framework termed Mathematical Architecture Design for Deep CNN (DeepMAD) is proposed to design high-performance CNN models in a principled way. In DeepMAD, a CNN network is modeled as an information processing system whose expressiveness and effectiveness can be analytically formulated by their structural parameters. Then a constrained mathematical programming (MP) problem is proposed to optimize these structural parameters. The MP problem can be easily solved by off-the-shelf MP solvers on CPUs with a small memory footprint. In addition, DeepMAD is a pure mathematical framework: no GPU or training data is required during network design. The superiority of DeepMAD is validated on multiple large-scale computer vision benchmark datasets. Notably on ImageNet-1k, only using conventional convolutional layers, DeepMAD achieves 0.7% and 1.5% higher top-1 accuracy than ConvNeXt and Swin on Tiny level, and 0.8% and 0.9% higher on Small level.
Over the past few years, the rapid development of deep learning technologies for computer vision has greatly promoted the performance of medical image segmentation (MedISeg). However, the recent MedISeg publications usually focus on presentations of the major contributions (e.g., network architectures, training strategies, and loss functions) while unwittingly ignoring some marginal implementation details (also known as "tricks"), leading to a potential problem of the unfair experimental result comparisons. In this paper, we collect a series of MedISeg tricks for different model implementation phases (i.e., pre-training model, data pre-processing, data augmentation, model implementation, model inference, and result post-processing), and experimentally explore the effectiveness of these tricks on the consistent baseline models. Compared to paper-driven surveys that only blandly focus on the advantages and limitation analyses of segmentation models, our work provides a large number of solid experiments and is more technically operable. With the extensive experimental results on both the representative 2D and 3D medical image datasets, we explicitly clarify the effect of these tricks. Moreover, based on the surveyed tricks, we also open-sourced a strong MedISeg repository, where each of its components has the advantage of plug-and-play. We believe that this milestone work not only completes a comprehensive and complementary survey of the state-of-the-art MedISeg approaches, but also offers a practical guide for addressing the future medical image processing challenges including but not limited to small dataset learning, class imbalance learning, multi-modality learning, and domain adaptation. The code has been released at: //github.com/hust-linyi/MedISeg
Image segmentation is a key topic in image processing and computer vision with applications such as scene understanding, medical image analysis, robotic perception, video surveillance, augmented reality, and image compression, among many others. Various algorithms for image segmentation have been developed in the literature. Recently, due to the success of deep learning models in a wide range of vision applications, there has been a substantial amount of works aimed at developing image segmentation approaches using deep learning models. In this survey, we provide a comprehensive review of the literature at the time of this writing, covering a broad spectrum of pioneering works for semantic and instance-level segmentation, including fully convolutional pixel-labeling networks, encoder-decoder architectures, multi-scale and pyramid based approaches, recurrent networks, visual attention models, and generative models in adversarial settings. We investigate the similarity, strengths and challenges of these deep learning models, examine the most widely used datasets, report performances, and discuss promising future research directions in this area.
Deep learning has become the most widely used approach for cardiac image segmentation in recent years. In this paper, we provide a review of over 100 cardiac image segmentation papers using deep learning, which covers common imaging modalities including magnetic resonance imaging (MRI), computed tomography (CT), and ultrasound (US) and major anatomical structures of interest (ventricles, atria and vessels). In addition, a summary of publicly available cardiac image datasets and code repositories are included to provide a base for encouraging reproducible research. Finally, we discuss the challenges and limitations with current deep learning-based approaches (scarcity of labels, model generalizability across different domains, interpretability) and suggest potential directions for future research.
In this paper, we adopt 3D Convolutional Neural Networks to segment volumetric medical images. Although deep neural networks have been proven to be very effective on many 2D vision tasks, it is still challenging to apply them to 3D tasks due to the limited amount of annotated 3D data and limited computational resources. We propose a novel 3D-based coarse-to-fine framework to effectively and efficiently tackle these challenges. The proposed 3D-based framework outperforms the 2D counterpart to a large margin since it can leverage the rich spatial infor- mation along all three axes. We conduct experiments on two datasets which include healthy and pathological pancreases respectively, and achieve the current state-of-the-art in terms of Dice-S{\o}rensen Coefficient (DSC). On the NIH pancreas segmentation dataset, we outperform the previous best by an average of over 2%, and the worst case is improved by 7% to reach almost 70%, which indicates the reliability of our framework in clinical applications.
Deep neural network architectures have traditionally been designed and explored with human expertise in a long-lasting trial-and-error process. This process requires huge amount of time, expertise, and resources. To address this tedious problem, we propose a novel algorithm to optimally find hyperparameters of a deep network architecture automatically. We specifically focus on designing neural architectures for medical image segmentation task. Our proposed method is based on a policy gradient reinforcement learning for which the reward function is assigned a segmentation evaluation utility (i.e., dice index). We show the efficacy of the proposed method with its low computational cost in comparison with the state-of-the-art medical image segmentation networks. We also present a new architecture design, a densely connected encoder-decoder CNN, as a strong baseline architecture to apply the proposed hyperparameter search algorithm. We apply the proposed algorithm to each layer of the baseline architectures. As an application, we train the proposed system on cine cardiac MR images from Automated Cardiac Diagnosis Challenge (ACDC) MICCAI 2017. Starting from a baseline segmentation architecture, the resulting network architecture obtains the state-of-the-art results in accuracy without performing any trial-and-error based architecture design approaches or close supervision of the hyperparameters changes.
Deep Convolutional Neural Networks have pushed the state-of-the art for semantic segmentation provided that a large amount of images together with pixel-wise annotations is available. Data collection is expensive and a solution to alleviate it is to use transfer learning. This reduces the amount of annotated data required for the network training but it does not get rid of this heavy processing step. We propose a method of transfer learning without annotations on the target task for datasets with redundant content and distinct pixel distributions. Our method takes advantage of the approximate content alignment of the images between two datasets when the approximation error prevents the reuse of annotation from one dataset to another. Given the annotations for only one dataset, we train a first network in a supervised manner. This network autonomously learns to generate deep data representations relevant to the semantic segmentation. Then the images in the new dataset, we train a new network to generate a deep data representation that matches the one from the first network on the previous dataset. The training consists in a regression between feature maps and does not require any annotations on the new dataset. We show that this method reaches performances similar to a classic transfer learning on the PASCAL VOC dataset with synthetic transformations.