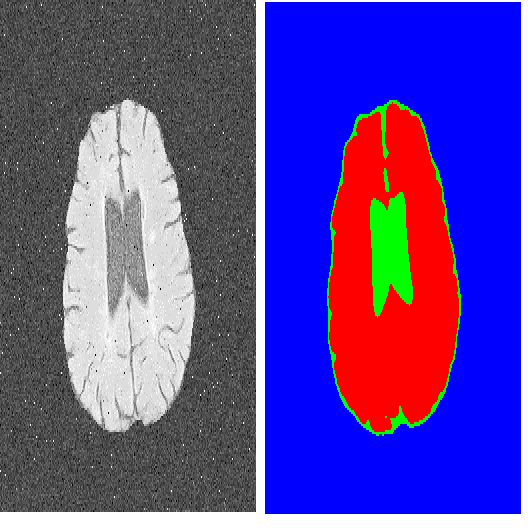
For medical image segmentation, contrastive learning is the dominant practice to improve the quality of visual representations by contrasting semantically similar and dissimilar pairs of samples. This is enabled by the observation that without accessing ground truth label, negative examples with truly dissimilar anatomical features, if sampled, can significantly improve the performance. In reality, however, these samples may come from similar anatomical features and the models may struggle to distinguish the minority tail-class samples, making the tail classes more prone to misclassification, both of which typically lead to model collapse. In this paper, we propose ARCO, a semi-supervised contrastive learning (CL) framework with stratified group sampling theory in medical image segmentation. In particular, we first propose building ARCO through the concept of variance-reduced estimation, and show that certain variance-reduction techniques are particularly beneficial in medical image segmentation tasks with extremely limited labels. Furthermore, we theoretically prove these sampling techniques are universal in variance reduction. Finally, we experimentally validate our approaches on three benchmark datasets with different label settings, and our methods consistently outperform state-of-the-art semi- and fully-supervised methods. Additionally, we augment the CL frameworks with these sampling techniques and demonstrate significant gains over previous methods. We believe our work is an important step towards semi-supervised medical image segmentation by quantifying the limitation of current self-supervision objectives for accomplishing medical image analysis tasks.
相關內容
How to ensure fairness is an important topic in federated learning (FL). Recent studies have investigated how to reward clients based on their contribution (collaboration fairness), and how to achieve uniformity of performance across clients (performance fairness). Despite achieving progress on either one, we argue that it is critical to consider them together, in order to engage and motivate more diverse clients joining FL to derive a high-quality global model. In this work, we propose a novel method to optimize both types of fairness simultaneously. Specifically, we propose to estimate client contribution in gradient and data space. In gradient space, we monitor the gradient direction differences of each client with respect to others. And in data space, we measure the prediction error on client data using an auxiliary model. Based on this contribution estimation, we propose a FL method, federated training via contribution estimation (FedCE), i.e., using estimation as global model aggregation weights. We have theoretically analyzed our method and empirically evaluated it on two real-world medical datasets. The effectiveness of our approach has been validated with significant performance improvements, better collaboration fairness, better performance fairness, and comprehensive analytical studies.
Semi-supervised medical image segmentation has attracted much attention in recent years because of the high cost of medical image annotations. In this paper, we propose a novel Inherent Consistent Learning (ICL) method, which aims to learn robust semantic category representations through the semantic consistency guidance of labeled and unlabeled data to help segmentation. In practice, we introduce two external modules namely Supervised Semantic Proxy Adaptor (SSPA) and Unsupervised Semantic Consistent Learner (USCL) that based on the attention mechanism to align the semantic category representations of labeled and unlabeled data, as well as update the global semantic representations over the entire training set. The proposed ICL is a plug-and-play scheme for various network architectures and the two modules are not involved in the testing stage. Experimental results on three public benchmarks show that the proposed method can outperform the state-of-the-art especially when the number of annotated data is extremely limited. Code is available at: //github.com/zhuye98/ICL.git.
Despite the significant progress that depth-based 3D hand pose estimation methods have made in recent years, they still require a large amount of labeled training data to achieve high accuracy. However, collecting such data is both costly and time-consuming. To tackle this issue, we propose a semi-supervised method to significantly reduce the dependence on labeled training data. The proposed method consists of two identical networks trained jointly: a teacher network and a student network. The teacher network is trained using both the available labeled and unlabeled samples. It leverages the unlabeled samples via a loss formulation that encourages estimation equivariance under a set of affine transformations. The student network is trained using the unlabeled samples with their pseudo-labels provided by the teacher network. For inference at test time, only the student network is used. Extensive experiments demonstrate that the proposed method outperforms the state-of-the-art semi-supervised methods by large margins.
Over the past few years, the rapid development of deep learning technologies for computer vision has greatly promoted the performance of medical image segmentation (MedISeg). However, the recent MedISeg publications usually focus on presentations of the major contributions (e.g., network architectures, training strategies, and loss functions) while unwittingly ignoring some marginal implementation details (also known as "tricks"), leading to a potential problem of the unfair experimental result comparisons. In this paper, we collect a series of MedISeg tricks for different model implementation phases (i.e., pre-training model, data pre-processing, data augmentation, model implementation, model inference, and result post-processing), and experimentally explore the effectiveness of these tricks on the consistent baseline models. Compared to paper-driven surveys that only blandly focus on the advantages and limitation analyses of segmentation models, our work provides a large number of solid experiments and is more technically operable. With the extensive experimental results on both the representative 2D and 3D medical image datasets, we explicitly clarify the effect of these tricks. Moreover, based on the surveyed tricks, we also open-sourced a strong MedISeg repository, where each of its components has the advantage of plug-and-play. We believe that this milestone work not only completes a comprehensive and complementary survey of the state-of-the-art MedISeg approaches, but also offers a practical guide for addressing the future medical image processing challenges including but not limited to small dataset learning, class imbalance learning, multi-modality learning, and domain adaptation. The code has been released at: //github.com/hust-linyi/MedISeg
A key requirement for the success of supervised deep learning is a large labeled dataset - a condition that is difficult to meet in medical image analysis. Self-supervised learning (SSL) can help in this regard by providing a strategy to pre-train a neural network with unlabeled data, followed by fine-tuning for a downstream task with limited annotations. Contrastive learning, a particular variant of SSL, is a powerful technique for learning image-level representations. In this work, we propose strategies for extending the contrastive learning framework for segmentation of volumetric medical images in the semi-supervised setting with limited annotations, by leveraging domain-specific and problem-specific cues. Specifically, we propose (1) novel contrasting strategies that leverage structural similarity across volumetric medical images (domain-specific cue) and (2) a local version of the contrastive loss to learn distinctive representations of local regions that are useful for per-pixel segmentation (problem-specific cue). We carry out an extensive evaluation on three Magnetic Resonance Imaging (MRI) datasets. In the limited annotation setting, the proposed method yields substantial improvements compared to other self-supervision and semi-supervised learning techniques. When combined with a simple data augmentation technique, the proposed method reaches within 8% of benchmark performance using only two labeled MRI volumes for training, corresponding to only 4% (for ACDC) of the training data used to train the benchmark.
Image segmentation is a key topic in image processing and computer vision with applications such as scene understanding, medical image analysis, robotic perception, video surveillance, augmented reality, and image compression, among many others. Various algorithms for image segmentation have been developed in the literature. Recently, due to the success of deep learning models in a wide range of vision applications, there has been a substantial amount of works aimed at developing image segmentation approaches using deep learning models. In this survey, we provide a comprehensive review of the literature at the time of this writing, covering a broad spectrum of pioneering works for semantic and instance-level segmentation, including fully convolutional pixel-labeling networks, encoder-decoder architectures, multi-scale and pyramid based approaches, recurrent networks, visual attention models, and generative models in adversarial settings. We investigate the similarity, strengths and challenges of these deep learning models, examine the most widely used datasets, report performances, and discuss promising future research directions in this area.
With the rapid increase of large-scale, real-world datasets, it becomes critical to address the problem of long-tailed data distribution (i.e., a few classes account for most of the data, while most classes are under-represented). Existing solutions typically adopt class re-balancing strategies such as re-sampling and re-weighting based on the number of observations for each class. In this work, we argue that as the number of samples increases, the additional benefit of a newly added data point will diminish. We introduce a novel theoretical framework to measure data overlap by associating with each sample a small neighboring region rather than a single point. The effective number of samples is defined as the volume of samples and can be calculated by a simple formula $(1-\beta^{n})/(1-\beta)$, where $n$ is the number of samples and $\beta \in [0,1)$ is a hyperparameter. We design a re-weighting scheme that uses the effective number of samples for each class to re-balance the loss, thereby yielding a class-balanced loss. Comprehensive experiments are conducted on artificially induced long-tailed CIFAR datasets and large-scale datasets including ImageNet and iNaturalist. Our results show that when trained with the proposed class-balanced loss, the network is able to achieve significant performance gains on long-tailed datasets.
The U-Net was presented in 2015. With its straight-forward and successful architecture it quickly evolved to a commonly used benchmark in medical image segmentation. The adaptation of the U-Net to novel problems, however, comprises several degrees of freedom regarding the exact architecture, preprocessing, training and inference. These choices are not independent of each other and substantially impact the overall performance. The present paper introduces the nnU-Net ('no-new-Net'), which refers to a robust and self-adapting framework on the basis of 2D and 3D vanilla U-Nets. We argue the strong case for taking away superfluous bells and whistles of many proposed network designs and instead focus on the remaining aspects that make out the performance and generalizability of a method. We evaluate the nnU-Net in the context of the Medical Segmentation Decathlon challenge, which measures segmentation performance in ten disciplines comprising distinct entities, image modalities, image geometries and dataset sizes, with no manual adjustments between datasets allowed. At the time of manuscript submission, nnU-Net achieves the highest mean dice scores across all classes and seven phase 1 tasks (except class 1 in BrainTumour) in the online leaderboard of the challenge.
Medical image segmentation requires consensus ground truth segmentations to be derived from multiple expert annotations. A novel approach is proposed that obtains consensus segmentations from experts using graph cuts (GC) and semi supervised learning (SSL). Popular approaches use iterative Expectation Maximization (EM) to estimate the final annotation and quantify annotator's performance. Such techniques pose the risk of getting trapped in local minima. We propose a self consistency (SC) score to quantify annotator consistency using low level image features. SSL is used to predict missing annotations by considering global features and local image consistency. The SC score also serves as the penalty cost in a second order Markov random field (MRF) cost function optimized using graph cuts to derive the final consensus label. Graph cut obtains a global maximum without an iterative procedure. Experimental results on synthetic images, real data of Crohn's disease patients and retinal images show our final segmentation to be accurate and more consistent than competing methods.

Recent advances in 3D fully convolutional networks (FCN) have made it feasible to produce dense voxel-wise predictions of volumetric images. In this work, we show that a multi-class 3D FCN trained on manually labeled CT scans of several anatomical structures (ranging from the large organs to thin vessels) can achieve competitive segmentation results, while avoiding the need for handcrafting features or training class-specific models. To this end, we propose a two-stage, coarse-to-fine approach that will first use a 3D FCN to roughly define a candidate region, which will then be used as input to a second 3D FCN. This reduces the number of voxels the second FCN has to classify to ~10% and allows it to focus on more detailed segmentation of the organs and vessels. We utilize training and validation sets consisting of 331 clinical CT images and test our models on a completely unseen data collection acquired at a different hospital that includes 150 CT scans, targeting three anatomical organs (liver, spleen, and pancreas). In challenging organs such as the pancreas, our cascaded approach improves the mean Dice score from 68.5 to 82.2%, achieving the highest reported average score on this dataset. We compare with a 2D FCN method on a separate dataset of 240 CT scans with 18 classes and achieve a significantly higher performance in small organs and vessels. Furthermore, we explore fine-tuning our models to different datasets. Our experiments illustrate the promise and robustness of current 3D FCN based semantic segmentation of medical images, achieving state-of-the-art results. Our code and trained models are available for download: //github.com/holgerroth/3Dunet_abdomen_cascade.