Accurate segmentation of large areas from very high spatial-resolution (VHR) remote sensing imagery remains a challenging issue in image analysis. Existing supervised and unsupervised methods both suffer from the large variance of object sizes and the difficulty in scale selection, which often result in poor segmentation accuracies. To address the above challenges, we propose a deep learning-based region-merging method (DeepMerge) to handle the segmentation in large VHR images by integrating a Transformer with a multi-level embedding module, a segment-based feature embedding module and a region-adjacency graph model. In addition, we propose a modified binary tree sampling method to generate multi-level inputs from initial segmentation results, serving as inputs for the DeepMerge model. To our best knowledge, the proposed method is the first to use deep learning to learn the similarity between adjacent segments for region-merging. The proposed DeepMerge method is validated using a remote sensing image of 0.55m resolution covering an area of 5,660 km^2 acquired from Google Earth. The experimental results show that the proposed DeepMerge with the highest F value (0.9446) and the lowest TE (0.0962) and ED2 (0.8989) is able to correctly segment objects of different sizes and outperforms all selected competing segmentation methods from both quantitative and qualitative assessments.
相關內容
Medical imaging has witnessed remarkable progress but usually requires a large amount of high-quality annotated data which is time-consuming and costly to obtain. To alleviate this burden, semi-supervised learning has garnered attention as a potential solution. In this paper, we present Meta-Learning for Bootstrapping Medical Image Segmentation (MLB-Seg), a novel method for tackling the challenge of semi-supervised medical image segmentation. Specifically, our approach first involves training a segmentation model on a small set of clean labeled images to generate initial labels for unlabeled data. To further optimize this bootstrapping process, we introduce a per-pixel weight mapping system that dynamically assigns weights to both the initialized labels and the model's own predictions. These weights are determined using a meta-process that prioritizes pixels with loss gradient directions closer to those of clean data, which is based on a small set of precisely annotated images. To facilitate the meta-learning process, we additionally introduce a consistency-based Pseudo Label Enhancement (PLE) scheme that improves the quality of the model's own predictions by ensembling predictions from various augmented versions of the same input. In order to improve the quality of the weight maps obtained through multiple augmentations of a single input, we introduce a mean teacher into the PLE scheme. This method helps to reduce noise in the weight maps and stabilize its generation process. Our extensive experimental results on public atrial and prostate segmentation datasets demonstrate that our proposed method achieves state-of-the-art results under semi-supervision. Our code is available at //github.com/aijinrjinr/MLB-Seg.
Unsupervised domain adaptation (UDA) has increasingly gained interests for its capacity to transfer the knowledge learned from a labeled source domain to an unlabeled target domain. However, typical UDA methods require concurrent access to both the source and target domain data, which largely limits its application in medical scenarios where source data is often unavailable due to privacy concern. To tackle the source data-absent problem, we present a novel two-stage source-free domain adaptation (SFDA) framework for medical image segmentation, where only a well-trained source segmentation model and unlabeled target data are available during domain adaptation. Specifically, in the prototype-anchored feature alignment stage, we first utilize the weights of the pre-trained pixel-wise classifier as source prototypes, which preserve the information of source features. Then, we introduce the bi-directional transport to align the target features with class prototypes by minimizing its expected cost. On top of that, a contrastive learning stage is further devised to utilize those pixels with unreliable predictions for a more compact target feature distribution. Extensive experiments on a cross-modality medical segmentation task demonstrate the superiority of our method in large domain discrepancy settings compared with the state-of-the-art SFDA approaches and even some UDA methods. Code is available at //github.com/CSCYQJ/MICCAI23-ProtoContra-SFDA.
Deep learning-based semi-supervised learning (SSL) algorithms have led to promising results in medical images segmentation and can alleviate doctors' expensive annotations by leveraging unlabeled data. However, most of the existing SSL algorithms in literature tend to regularize the model training by perturbing networks and/or data. Observing that multi/dual-task learning attends to various levels of information which have inherent prediction perturbation, we ask the question in this work: can we explicitly build task-level regularization rather than implicitly constructing networks- and/or data-level perturbation-and-transformation for SSL? To answer this question, we propose a novel dual-task-consistency semi-supervised framework for the first time. Concretely, we use a dual-task deep network that jointly predicts a pixel-wise segmentation map and a geometry-aware level set representation of the target. The level set representation is converted to an approximated segmentation map through a differentiable task transform layer. Simultaneously, we introduce a dual-task consistency regularization between the level set-derived segmentation maps and directly predicted segmentation maps for both labeled and unlabeled data. Extensive experiments on two public datasets show that our method can largely improve the performance by incorporating the unlabeled data. Meanwhile, our framework outperforms the state-of-the-art semi-supervised medical image segmentation methods. Code is available at: //github.com/Luoxd1996/DTC
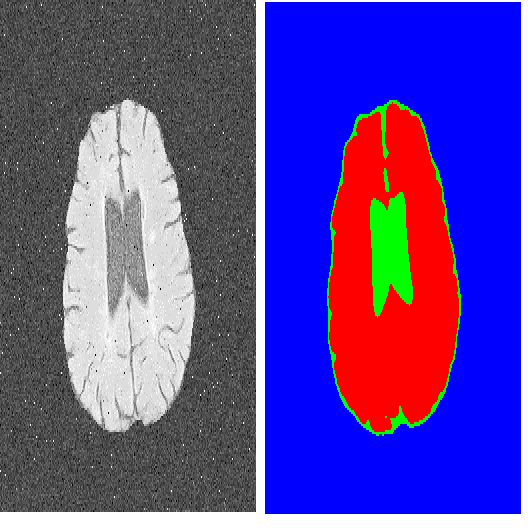
A key requirement for the success of supervised deep learning is a large labeled dataset - a condition that is difficult to meet in medical image analysis. Self-supervised learning (SSL) can help in this regard by providing a strategy to pre-train a neural network with unlabeled data, followed by fine-tuning for a downstream task with limited annotations. Contrastive learning, a particular variant of SSL, is a powerful technique for learning image-level representations. In this work, we propose strategies for extending the contrastive learning framework for segmentation of volumetric medical images in the semi-supervised setting with limited annotations, by leveraging domain-specific and problem-specific cues. Specifically, we propose (1) novel contrasting strategies that leverage structural similarity across volumetric medical images (domain-specific cue) and (2) a local version of the contrastive loss to learn distinctive representations of local regions that are useful for per-pixel segmentation (problem-specific cue). We carry out an extensive evaluation on three Magnetic Resonance Imaging (MRI) datasets. In the limited annotation setting, the proposed method yields substantial improvements compared to other self-supervision and semi-supervised learning techniques. When combined with a simple data augmentation technique, the proposed method reaches within 8% of benchmark performance using only two labeled MRI volumes for training, corresponding to only 4% (for ACDC) of the training data used to train the benchmark.
Deep learning has become the most widely used approach for cardiac image segmentation in recent years. In this paper, we provide a review of over 100 cardiac image segmentation papers using deep learning, which covers common imaging modalities including magnetic resonance imaging (MRI), computed tomography (CT), and ultrasound (US) and major anatomical structures of interest (ventricles, atria and vessels). In addition, a summary of publicly available cardiac image datasets and code repositories are included to provide a base for encouraging reproducible research. Finally, we discuss the challenges and limitations with current deep learning-based approaches (scarcity of labels, model generalizability across different domains, interpretability) and suggest potential directions for future research.
It is always well believed that modeling relationships between objects would be helpful for representing and eventually describing an image. Nevertheless, there has not been evidence in support of the idea on image description generation. In this paper, we introduce a new design to explore the connections between objects for image captioning under the umbrella of attention-based encoder-decoder framework. Specifically, we present Graph Convolutional Networks plus Long Short-Term Memory (dubbed as GCN-LSTM) architecture that novelly integrates both semantic and spatial object relationships into image encoder. Technically, we build graphs over the detected objects in an image based on their spatial and semantic connections. The representations of each region proposed on objects are then refined by leveraging graph structure through GCN. With the learnt region-level features, our GCN-LSTM capitalizes on LSTM-based captioning framework with attention mechanism for sentence generation. Extensive experiments are conducted on COCO image captioning dataset, and superior results are reported when comparing to state-of-the-art approaches. More remarkably, GCN-LSTM increases CIDEr-D performance from 120.1% to 128.7% on COCO testing set.
In this paper, we focus on three problems in deep learning based medical image segmentation. Firstly, U-net, as a popular model for medical image segmentation, is difficult to train when convolutional layers increase even though a deeper network usually has a better generalization ability because of more learnable parameters. Secondly, the exponential ReLU (ELU), as an alternative of ReLU, is not much different from ReLU when the network of interest gets deep. Thirdly, the Dice loss, as one of the pervasive loss functions for medical image segmentation, is not effective when the prediction is close to ground truth and will cause oscillation during training. To address the aforementioned three problems, we propose and validate a deeper network that can fit medical image datasets that are usually small in the sample size. Meanwhile, we propose a new loss function to accelerate the learning process and a combination of different activation functions to improve the network performance. Our experimental results suggest that our network is comparable or superior to state-of-the-art methods.
Medical image segmentation requires consensus ground truth segmentations to be derived from multiple expert annotations. A novel approach is proposed that obtains consensus segmentations from experts using graph cuts (GC) and semi supervised learning (SSL). Popular approaches use iterative Expectation Maximization (EM) to estimate the final annotation and quantify annotator's performance. Such techniques pose the risk of getting trapped in local minima. We propose a self consistency (SC) score to quantify annotator consistency using low level image features. SSL is used to predict missing annotations by considering global features and local image consistency. The SC score also serves as the penalty cost in a second order Markov random field (MRF) cost function optimized using graph cuts to derive the final consensus label. Graph cut obtains a global maximum without an iterative procedure. Experimental results on synthetic images, real data of Crohn's disease patients and retinal images show our final segmentation to be accurate and more consistent than competing methods.
Deep Convolutional Neural Networks have pushed the state-of-the art for semantic segmentation provided that a large amount of images together with pixel-wise annotations is available. Data collection is expensive and a solution to alleviate it is to use transfer learning. This reduces the amount of annotated data required for the network training but it does not get rid of this heavy processing step. We propose a method of transfer learning without annotations on the target task for datasets with redundant content and distinct pixel distributions. Our method takes advantage of the approximate content alignment of the images between two datasets when the approximation error prevents the reuse of annotation from one dataset to another. Given the annotations for only one dataset, we train a first network in a supervised manner. This network autonomously learns to generate deep data representations relevant to the semantic segmentation. Then the images in the new dataset, we train a new network to generate a deep data representation that matches the one from the first network on the previous dataset. The training consists in a regression between feature maps and does not require any annotations on the new dataset. We show that this method reaches performances similar to a classic transfer learning on the PASCAL VOC dataset with synthetic transformations.
Convolutional networks (ConvNets) have achieved great successes in various challenging vision tasks. However, the performance of ConvNets would degrade when encountering the domain shift. The domain adaptation is more significant while challenging in the field of biomedical image analysis, where cross-modality data have largely different distributions. Given that annotating the medical data is especially expensive, the supervised transfer learning approaches are not quite optimal. In this paper, we propose an unsupervised domain adaptation framework with adversarial learning for cross-modality biomedical image segmentations. Specifically, our model is based on a dilated fully convolutional network for pixel-wise prediction. Moreover, we build a plug-and-play domain adaptation module (DAM) to map the target input to features which are aligned with source domain feature space. A domain critic module (DCM) is set up for discriminating the feature space of both domains. We optimize the DAM and DCM via an adversarial loss without using any target domain label. Our proposed method is validated by adapting a ConvNet trained with MRI images to unpaired CT data for cardiac structures segmentations, and achieved very promising results.