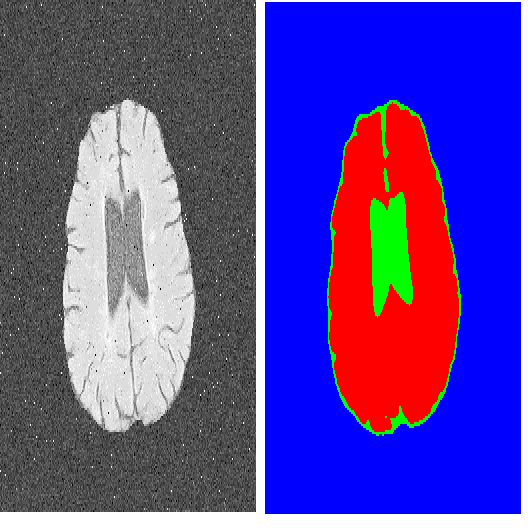
Although the preservation of shape continuity and physiological anatomy is a natural assumption in the segmentation of medical images, it is often neglected by deep learning methods that mostly aim for the statistical modeling of input data as pixels rather than interconnected structures. In biological structures, however, organs are not separate entities; for example, in reality, a severed vessel is an indication of an underlying problem, but traditional segmentation models are not designed to strictly enforce the continuity of anatomy, potentially leading to inaccurate medical diagnoses. To address this issue, we propose a graph-based approach that enforces the continuity and connectivity of anatomical topology in medical images. Our method encodes the continuity of shapes as a graph constraint, ensuring that the network's predictions maintain this continuity. We evaluate our method on two public benchmarks on retinal vessel segmentation, showing significant improvements in connectivity metrics compared to traditional methods while getting better or on-par performance on segmentation metrics.
相關內容
Source:
We study the problem of model selection in causal inference, specifically for the case of conditional average treatment effect (CATE) estimation under binary treatments. Unlike model selection in machine learning, there is no perfect analogue of cross-validation as we do not observe the counterfactual potential outcome for any data point. Towards this, there have been a variety of proxy metrics proposed in the literature, that depend on auxiliary nuisance models estimated from the observed data (propensity score model, outcome regression model). However, the effectiveness of these metrics has only been studied on synthetic datasets as we can access the counterfactual data for them. We conduct an extensive empirical analysis to judge the performance of these metrics introduced in the literature, and novel ones introduced in this work, where we utilize the latest advances in generative modeling to incorporate multiple realistic datasets. Our analysis suggests novel model selection strategies based on careful hyperparameter tuning of CATE estimators and causal ensembling.
When estimating a regression model, we might have data where some labels are missing, or our data might be biased by a selection mechanism. When the response or selection mechanism is ignorable (i.e., independent of the response variable given the features) one can use off-the-shelf regression methods; in the nonignorable case one typically has to adjust for bias. We observe that privileged information (i.e. information that is only available during training) might render a nonignorable selection mechanism ignorable, and we refer to this scenario as Privilegedly Missing at Random (PMAR). We propose a novel imputation-based regression method, named repeated regression, that is suitable for PMAR. We also consider an importance weighted regression method, and a doubly robust combination of the two. The proposed methods are easy to implement with most popular out-of-the-box regression algorithms. We empirically assess the performance of the proposed methods with extensive simulated experiments and on a synthetically augmented real-world dataset. We conclude that repeated regression can appropriately correct for bias, and can have considerable advantage over weighted regression, especially when extrapolating to regions of the feature space where response is never observed.
In this paper, we present a notion of differential privacy (DP) for data that comes from different classes. Here, the class-membership is private information that needs to be protected. The proposed method is an output perturbation mechanism that adds noise to the release of query response such that the analyst is unable to infer the underlying class-label. The proposed DP method is capable of not only protecting the privacy of class-based data but also meets quality metrics of accuracy and is computationally efficient and practical. We illustrate the efficacy of the proposed method empirically while outperforming the baseline additive Gaussian noise mechanism. We also examine a real-world application and apply the proposed DP method to the autoregression and moving average (ARMA) forecasting method, protecting the privacy of the underlying data source. Case studies on the real-world advanced metering infrastructure (AMI) measurements of household power consumption validate the excellent performance of the proposed DP method while also satisfying the accuracy of forecasted power consumption measurements.
Deep learning-based semi-supervised learning (SSL) algorithms have led to promising results in medical images segmentation and can alleviate doctors' expensive annotations by leveraging unlabeled data. However, most of the existing SSL algorithms in literature tend to regularize the model training by perturbing networks and/or data. Observing that multi/dual-task learning attends to various levels of information which have inherent prediction perturbation, we ask the question in this work: can we explicitly build task-level regularization rather than implicitly constructing networks- and/or data-level perturbation-and-transformation for SSL? To answer this question, we propose a novel dual-task-consistency semi-supervised framework for the first time. Concretely, we use a dual-task deep network that jointly predicts a pixel-wise segmentation map and a geometry-aware level set representation of the target. The level set representation is converted to an approximated segmentation map through a differentiable task transform layer. Simultaneously, we introduce a dual-task consistency regularization between the level set-derived segmentation maps and directly predicted segmentation maps for both labeled and unlabeled data. Extensive experiments on two public datasets show that our method can largely improve the performance by incorporating the unlabeled data. Meanwhile, our framework outperforms the state-of-the-art semi-supervised medical image segmentation methods. Code is available at: //github.com/Luoxd1996/DTC
Graph Neural Networks (GNNs), which generalize deep neural networks to graph-structured data, have drawn considerable attention and achieved state-of-the-art performance in numerous graph related tasks. However, existing GNN models mainly focus on designing graph convolution operations. The graph pooling (or downsampling) operations, that play an important role in learning hierarchical representations, are usually overlooked. In this paper, we propose a novel graph pooling operator, called Hierarchical Graph Pooling with Structure Learning (HGP-SL), which can be integrated into various graph neural network architectures. HGP-SL incorporates graph pooling and structure learning into a unified module to generate hierarchical representations of graphs. More specifically, the graph pooling operation adaptively selects a subset of nodes to form an induced subgraph for the subsequent layers. To preserve the integrity of graph's topological information, we further introduce a structure learning mechanism to learn a refined graph structure for the pooled graph at each layer. By combining HGP-SL operator with graph neural networks, we perform graph level representation learning with focus on graph classification task. Experimental results on six widely used benchmarks demonstrate the effectiveness of our proposed model.
In this paper, we present an accurate and scalable approach to the face clustering task. We aim at grouping a set of faces by their potential identities. We formulate this task as a link prediction problem: a link exists between two faces if they are of the same identity. The key idea is that we find the local context in the feature space around an instance (face) contains rich information about the linkage relationship between this instance and its neighbors. By constructing sub-graphs around each instance as input data, which depict the local context, we utilize the graph convolution network (GCN) to perform reasoning and infer the likelihood of linkage between pairs in the sub-graphs. Experiments show that our method is more robust to the complex distribution of faces than conventional methods, yielding favorably comparable results to state-of-the-art methods on standard face clustering benchmarks, and is scalable to large datasets. Furthermore, we show that the proposed method does not need the number of clusters as prior, is aware of noises and outliers, and can be extended to a multi-view version for more accurate clustering accuracy.
In this paper, we focus on three problems in deep learning based medical image segmentation. Firstly, U-net, as a popular model for medical image segmentation, is difficult to train when convolutional layers increase even though a deeper network usually has a better generalization ability because of more learnable parameters. Secondly, the exponential ReLU (ELU), as an alternative of ReLU, is not much different from ReLU when the network of interest gets deep. Thirdly, the Dice loss, as one of the pervasive loss functions for medical image segmentation, is not effective when the prediction is close to ground truth and will cause oscillation during training. To address the aforementioned three problems, we propose and validate a deeper network that can fit medical image datasets that are usually small in the sample size. Meanwhile, we propose a new loss function to accelerate the learning process and a combination of different activation functions to improve the network performance. Our experimental results suggest that our network is comparable or superior to state-of-the-art methods.
A variety of deep neural networks have been applied in medical image segmentation and achieve good performance. Unlike natural images, medical images of the same imaging modality are characterized by the same pattern, which indicates that same normal organs or tissues locate at similar positions in the images. Thus, in this paper we try to incorporate the prior knowledge of medical images into the structure of neural networks such that the prior knowledge can be utilized for accurate segmentation. Based on this idea, we propose a novel deep network called knowledge-based fully convolutional network (KFCN) for medical image segmentation. The segmentation function and corresponding error is analyzed. We show the existence of an asymptotically stable region for KFCN which traditional FCN doesn't possess. Experiments validate our knowledge assumption about the incorporation of prior knowledge into the convolution kernels of KFCN and show that KFCN can achieve a reasonable segmentation and a satisfactory accuracy.
Medical image segmentation requires consensus ground truth segmentations to be derived from multiple expert annotations. A novel approach is proposed that obtains consensus segmentations from experts using graph cuts (GC) and semi supervised learning (SSL). Popular approaches use iterative Expectation Maximization (EM) to estimate the final annotation and quantify annotator's performance. Such techniques pose the risk of getting trapped in local minima. We propose a self consistency (SC) score to quantify annotator consistency using low level image features. SSL is used to predict missing annotations by considering global features and local image consistency. The SC score also serves as the penalty cost in a second order Markov random field (MRF) cost function optimized using graph cuts to derive the final consensus label. Graph cut obtains a global maximum without an iterative procedure. Experimental results on synthetic images, real data of Crohn's disease patients and retinal images show our final segmentation to be accurate and more consistent than competing methods.

Recent advances in 3D fully convolutional networks (FCN) have made it feasible to produce dense voxel-wise predictions of volumetric images. In this work, we show that a multi-class 3D FCN trained on manually labeled CT scans of several anatomical structures (ranging from the large organs to thin vessels) can achieve competitive segmentation results, while avoiding the need for handcrafting features or training class-specific models. To this end, we propose a two-stage, coarse-to-fine approach that will first use a 3D FCN to roughly define a candidate region, which will then be used as input to a second 3D FCN. This reduces the number of voxels the second FCN has to classify to ~10% and allows it to focus on more detailed segmentation of the organs and vessels. We utilize training and validation sets consisting of 331 clinical CT images and test our models on a completely unseen data collection acquired at a different hospital that includes 150 CT scans, targeting three anatomical organs (liver, spleen, and pancreas). In challenging organs such as the pancreas, our cascaded approach improves the mean Dice score from 68.5 to 82.2%, achieving the highest reported average score on this dataset. We compare with a 2D FCN method on a separate dataset of 240 CT scans with 18 classes and achieve a significantly higher performance in small organs and vessels. Furthermore, we explore fine-tuning our models to different datasets. Our experiments illustrate the promise and robustness of current 3D FCN based semantic segmentation of medical images, achieving state-of-the-art results. Our code and trained models are available for download: //github.com/holgerroth/3Dunet_abdomen_cascade.